|
eLinguistics.net's Mission & Statement:
-> Making language relatedness easily perceivable with a simple quantification. -> Setting up a 100% automated language classification. NEW!!!! -> Sinitic update, also in the GLOBAL LANGUAGE TREE! (April 2025) -> Updated LANGUAGE EVOLUTION TIMELINES! (Feb. 2024) This blog presents a completely computerized model for comparative linguistics. The quantification of language relationships is based on a probabilistic model, assessing basic vocabulary according to clear rules. It makes it possible to generate a 100% automated language classification into families and subfamilies and to generate objective distance values for language comparisons. You can compare languages in the calculator and get values for the relatedness (as a distance value) between 550 languages. An evolutionary tree summarizes all the results, showing how these languages relate to each other (single isolated languages are not displayed in the tree). This comparative linguistics approach takes you to a short digital trip in the history of languages... You will see how 18 words (when carefully chosen) can deliver values which are enough to calculate a distance between two and more languages and represent it on a tree. The distances are expressed as values between 0 (the nearest distance - so the same language) to 100 (biggest possible distance). Play with these values in the calculator! You will recognize proximities you can feel by yourself if you know some of the languages used in this study... |





|
A few examples to illustrate the idea behind this comparative linguistics project: the system's assessment for the distance from 0 to 100 between following languages is:
This gives you a first idea what this site is about. With the few examples above, you can conclude that the degree of relatedness between Russian and German (both Indo-European languages) is quite the same as the degree of relatedness between Finnish and Hungarian (both Finno-Ugric). Once we can get such values, we can generate a matrix. like this one, summing up distances between some languages (values from the few examples above have a green background in the matrix): 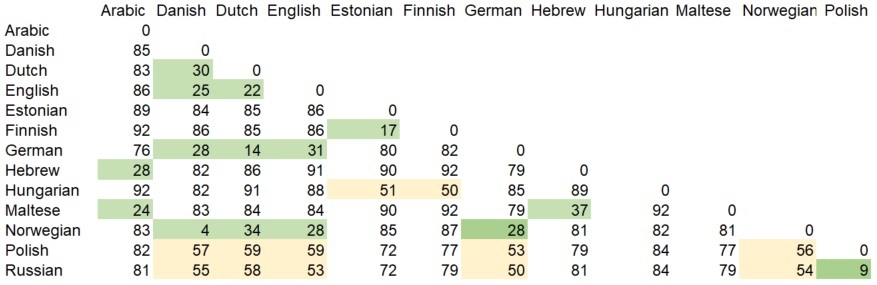
...and finally, out of this distance matrix, we generate an evolutionary tree - using the same system - and in fact the same software - like in genetics and biology (details under Resources): 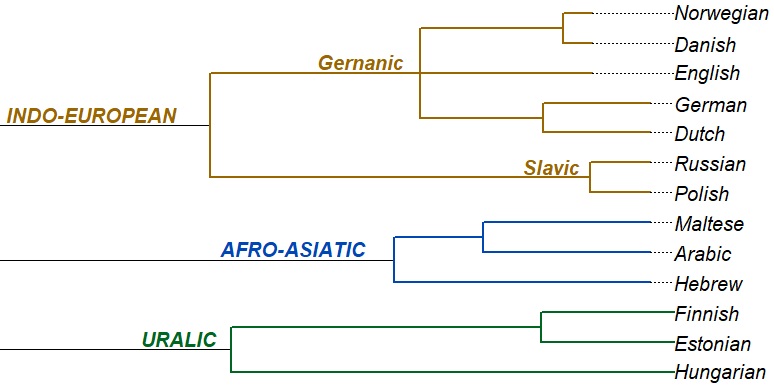
| |
Blog author: Vincent Beaufils - LinkedIn - Contact